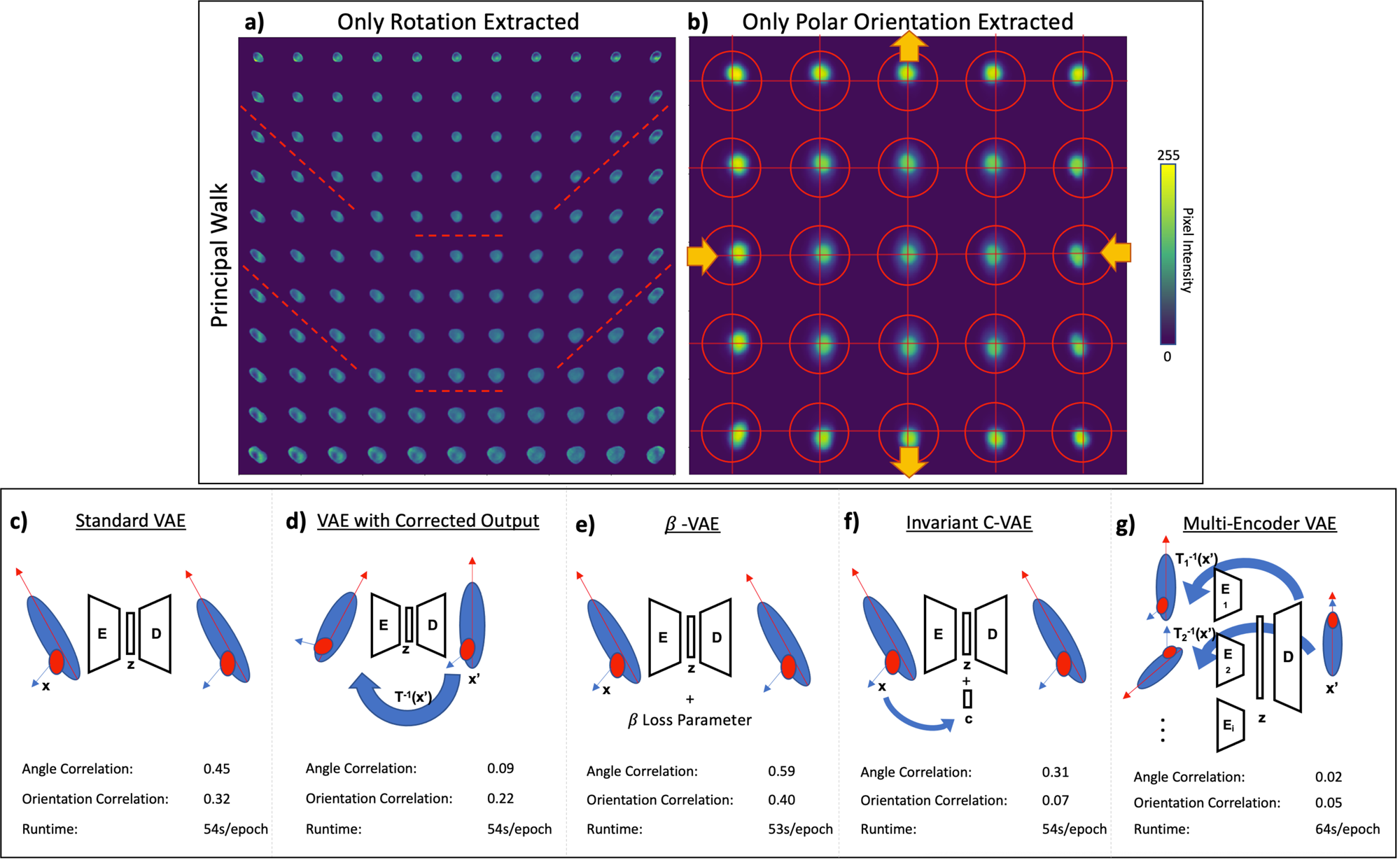
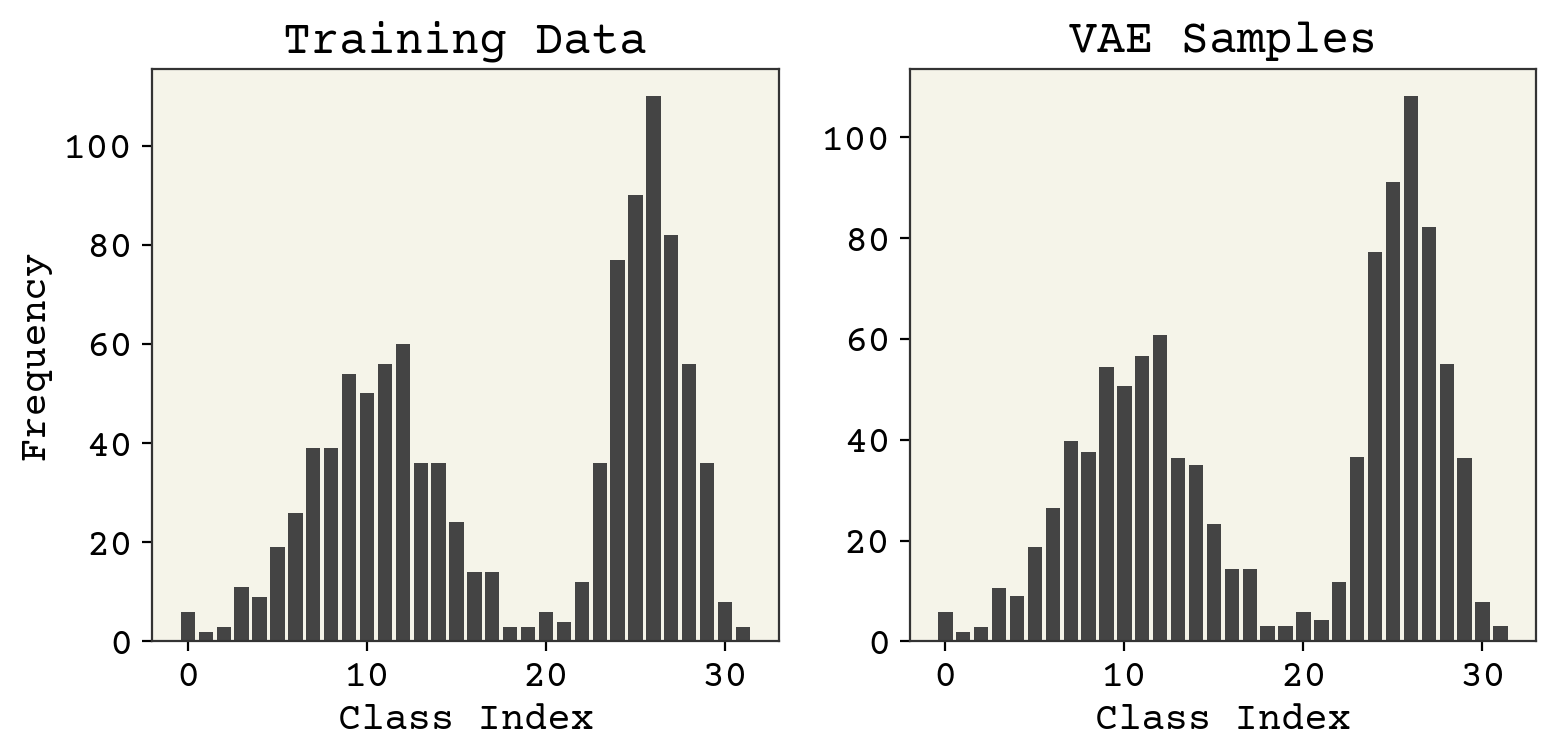
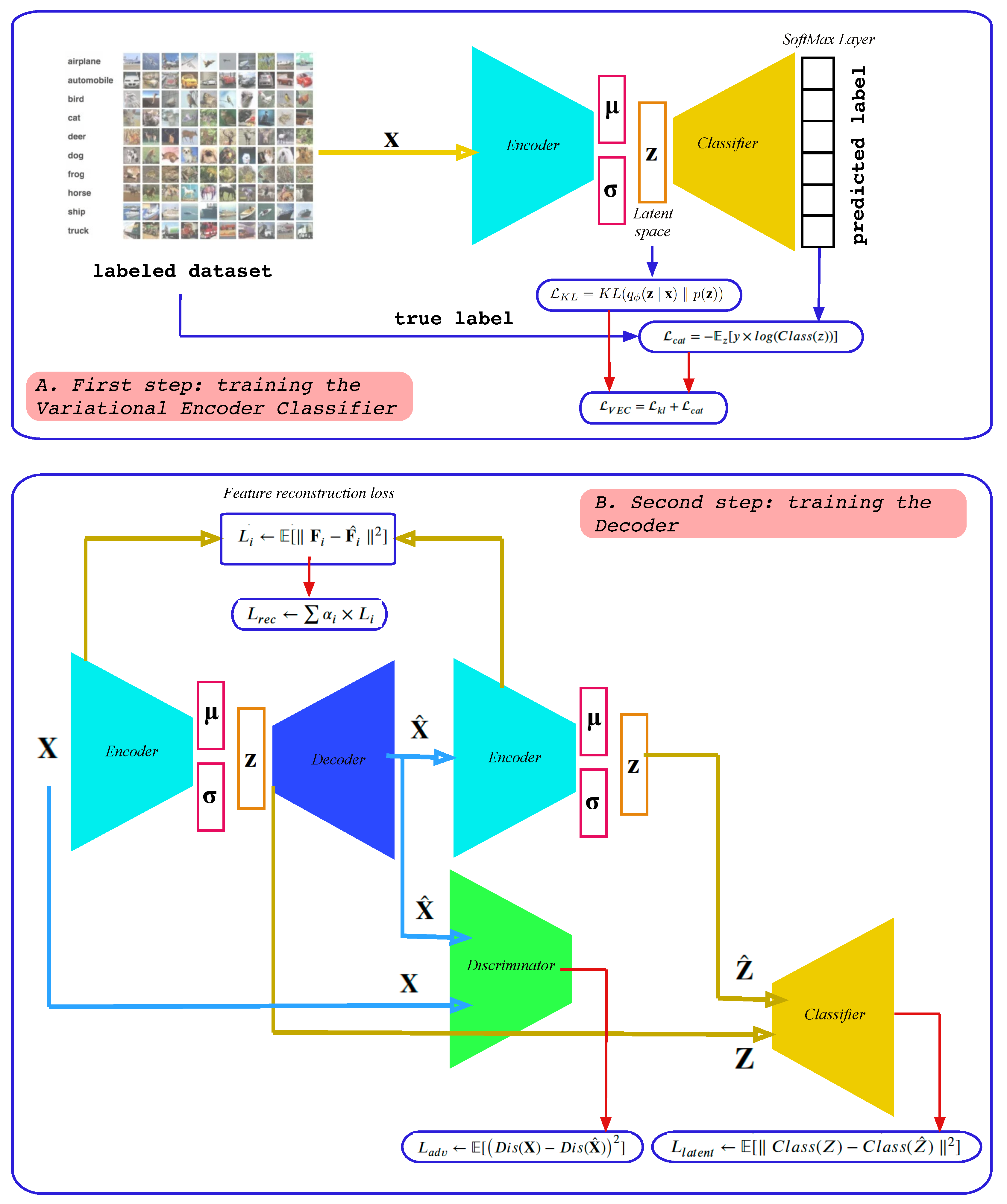
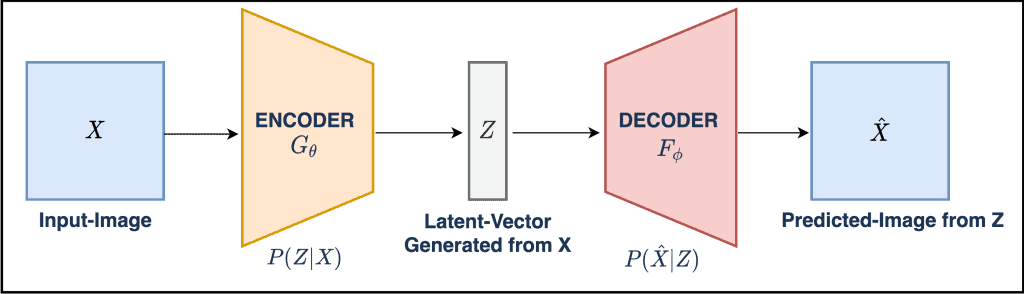
Partitioning variability in animal behavioral videos using semi-supervised variational autoencoders | PLOS Computational Biology

Quantitative comparison of principal component analysis and unsupervised deep learning using variational autoencoders for shape analysis of motile cells | bioRxiv

Encoding and exploring latent design space of optimal material structures via a VAE-LSTM model - ScienceDirect
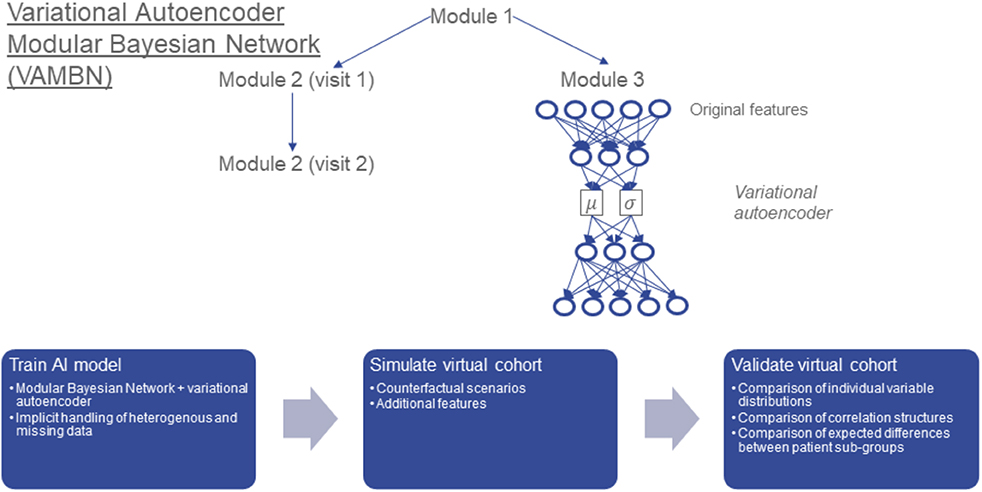
Frontiers | Variational Autoencoder Modular Bayesian Networks for Simulation of Heterogeneous Clinical Study Data

Variational autoencoder-based estimation of chronological age and changes in morphological features of teeth | Scientific Reports
Ig-VAE: Generative modeling of protein structure by direct 3D coordinate generation | PLOS Computational Biology

Molecules | Free Full-Text | VAE-Sim: A Novel Molecular Similarity Measure Based on a Variational Autoencoder
Predicting drug polypharmacology from cell morphology readouts using variational autoencoder latent space arithmetic | PLOS Computational Biology

Molecules | Free Full-Text | VAE-Sim: A Novel Molecular Similarity Measure Based on a Variational Autoencoder

Embedding of Molecular Structure Using Molecular Hypergraph Variational Autoencoder with Metric Learning - Koge - 2021 - Molecular Informatics - Wiley Online Library

L4 Latent Variable Models (VAE) -- CS294-158-SP20 Deep Unsupervised Learning -- UC Berkeley - YouTube

Learning Efficient, Collective Monte Carlo Moves with Variational Autoencoders | Journal of Chemical Theory and Computation